NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
JBrowse 2 Tutorial PAG 2022
This is the JBrowse 2 tutorial given at PAG 2022 via zoom. A recording of the tutorial is available at YouTube: https://youtu.be/rYqTYcZ56xs.
This tutorial covers the JBrowse Desktop application, including adding reference sequences, synteny data and views, annotation tracks and tracks with data from other sources. It finishes with examples of how to write JEXL to dynamically change the what tracks behave.
For JBrowse Web, see https://jbrowse.org/jb2/docs/superquickstart_web/ for a quick start to loading synteny tracks
Contents
Prerequisites
JBrowse 2 is both a desktop and server application. In this tutorial, we will focus on the desktop application to make our lives easier, but the server application is pretty easy to set up and has simple prerequisites.
Download and install
While we've installed JBrowse 2 on the conference computers (or we would have if we were there in person), if you'd like to follow along on your own computer, you can go to https://jbrowse.org/jb2/download/ and get the download for your platform and install it. It shouldn't take very long.
JBrowse Introduction
How and why JBrowse 2 is different from most other web-based genome browsers, including JBrowse and GBrowse.
Intro to JBrowse 2 (Google Slides)
Setting up JBrowse
Loading sequence
After installing JBrowse 2, open it using your operating systems preferred method, and you'll be greeted with a splash screen that has on part of it this dialog to open a new sequence:
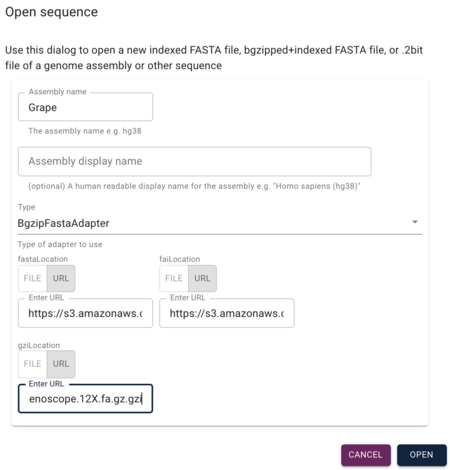
JBrowse supports a variety of forms of sequence data including "vanilla" FASTA, but for this example, we are going to use gzipped and faidx (FASTA indexed) files. To load those up, we'll use the grape FASTA file and it's indexes (ie, 'fai' and 'gzi' files):
https://s3.amazonaws.com/jbrowse.org/genomes/grape/Vvinifera_145_Genoscope.12X.fa.gz https://s3.amazonaws.com/jbrowse.org/genomes/grape/Vvinifera_145_Genoscope.12X.fa.gz.fai https://s3.amazonaws.com/jbrowse.org/genomes/grape/Vvinifera_145_Genoscope.12X.fa.gz.gzi
In the Open Sequence dialog, give the assembly a name (something creative, like "grape") and select BgzipFastaAdapter from the "type" menu, and then copy and paste the above URLs into the appropriate textfields under the "type" menu.
If we were creating a "normal" genome browser, we'd be done with adding sequence, but since we'd like to compare, we will also add the bgzipped and indexed FASTA file for peach. When we clicked on the "open sequence" button before, we were presented with a menu asking us what type of view we'd like, but first we have to add a second genome. What we need is in the Tools menu. Select "Open assembly manager," where you'll get a dialog that was very similar to what we used for grape. This time, we'll load the peach genome, so do the same things as before, and use these URLs:
https://s3.amazonaws.com/jbrowse.org/genomes/peach/Ppersica_298_v2.0.fa.gz https://s3.amazonaws.com/jbrowse.org/genomes/peach/Ppersica_298_v2.0.fa.gz.fai https://s3.amazonaws.com/jbrowse.org/genomes/peach/Ppersica_298_v2.0.fa.gz.gzi
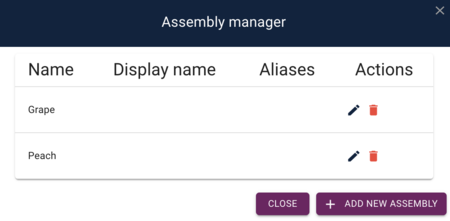
After adding the peach genome, we'll get a dialog that shows us that we have both genomes:
A note about preparing FASTA files
While we've provided compressed FASTA files, if you wanted to do this yourself, you would need bgzip from the Samtools package. The commands to create these files are:
bgzip -i yourfile.fa samtools faidx yourfile.fa.gz
Creating a comparative view
Now we'd like to create a comparison view. JBrowse 2 supports a few comparative views, but we'll start with a whole genome dotplot. For showing areas of synteny, we have a PAF file that looks like this:
Pp01 47851208 1388059 1391133 + chr8 22385789 1539799 1542834 703 3099 21 tp:A:P cm:i:73 s1:i:686 s2:i:439 dv:f:0.1377 rl:i:921840 Pp01 47851208 19134590 19135964 - chr15 20304914 6572992 6574378 659 1387 1 tp:A:P cm:i:85 s1:i:657 s2:i:638 dv:f:0.0768 rl:i:921840 Pp01 47851208 19134614 19135805 + chr17 17126926 16801080 16802270 638 1192 0 tp:A:S cm:i:79 s1:i:638 dv:f:0.0727 rl:i:921840 Pp01 47851208 43719774 43728648 - chr18 29360087 6242566 6251482 642 8964 54 tp:A:P cm:i:55 s1:i:620 s2:i:40 dv:f:0.2275 rl:i:921840 Pp01 47851208 40987755 40994103 + chr18 29360087 2664522 2670983 639 6461 51 tp:A:P cm:i:64 s1:i:620 s2:i:77 dv:f:0.1931 rl:i:921840 Pp01 47851208 19134590 19135968 - chr5 25021643 19591018 19592393 572 1379 0 tp:A:S cm:i:69 s1:i:572 dv:f:0.0910 rl:i:921840
PAF is a fairly simple file format relating two areas in genome coordinates. The PAF file was created with minimap2 like this:
minimap2 Vvinifera_457_Genoscope.12X.fa.gz Ppersica_298_v2.0.fa > Vvinifera_457_Genomescope.12X_vs_Ppersica_298_v2.0.paf
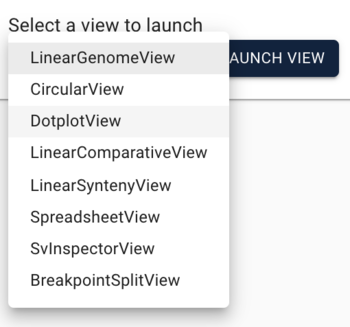
To load the peach-grape PAF, select "DotplotView" from the "Select a view to launch" menu.
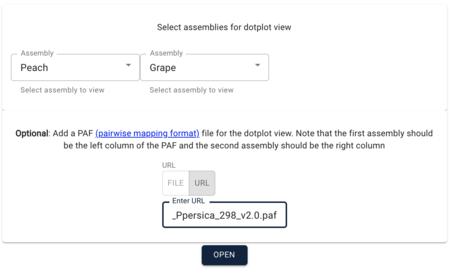
In the resulting dialog box, select Peach and then Grape for the assemblies to view. IMPORTANT: order here matters! Because the PAF file has the peach coordinates first, you have to use it first in this dialog box. After selecting the two assemblies, copy and paste this URL for the PAF file in to the optional PAF URL textfield:
https://s3.amazonaws.com/jbrowse.org/genomes/synteny/Vvinifera_457_Genomescope.12X_vs_Ppersica_298_v2.0.paf
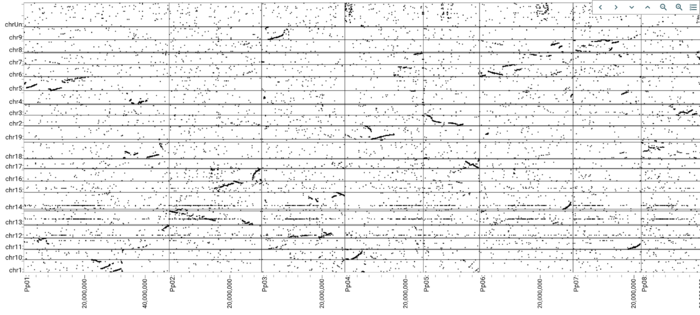
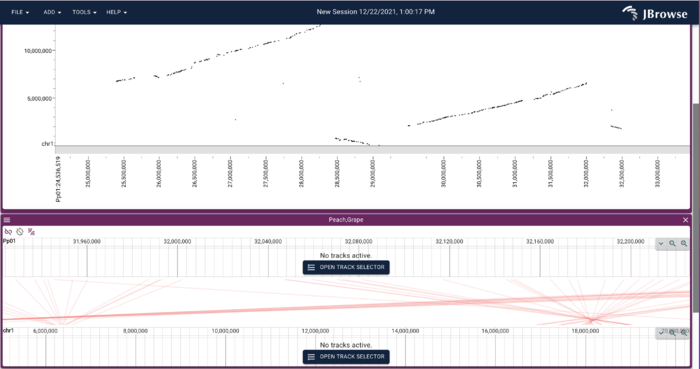
After clicking "Open", you get a dotplot that looks like this:
And of course, this isn't just an image, it is a genome-browsable interface, that you can click and drag to zoom into an region you like, even across multiple chromosomes.
Creating the synteny view
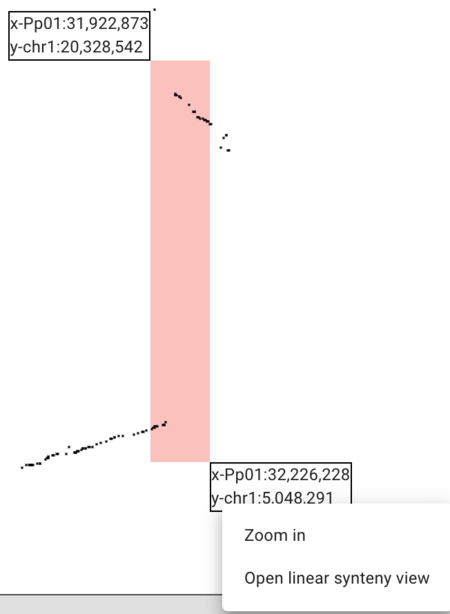
When we click and drag to make a rectangle, we get a popup menu asking whether we want to zoom in or open a synteny view. We can use this functionality to zoom in on a region we are interested in and then when we're happy with the region, we can click on the Open linear syntenic view option.
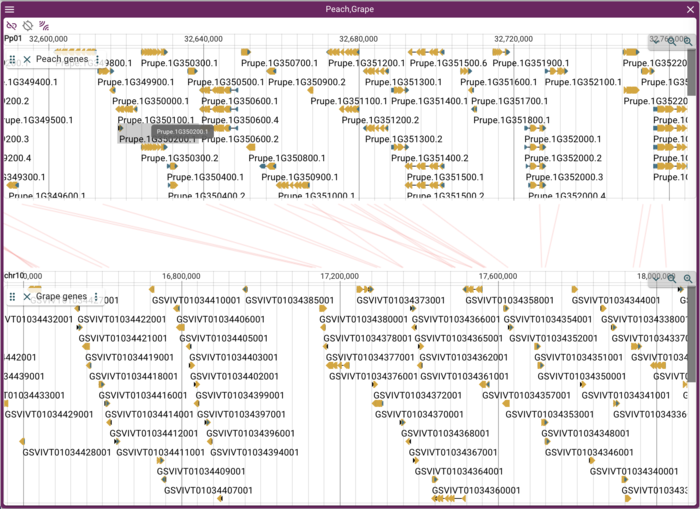
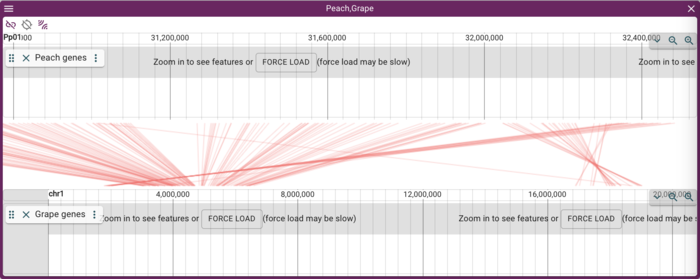
and the resulting syntenic view:
Adding gene annotations
This is nice--it shows lines or trapezoids of synteny, but is perhaps not as informative as it could be. The individual genome frames in the synteny view support adding other tracks (though if you add a lot, you better have a tall monitor), so we can add gene annotations. As it happens, we have gene annotation track data from a JBrowse 1 instance for both peach and grape (which was originally used for the GBrowse_syn tutorial), so we can add those. Note that this procedure will work for just about any sort of data file that we might might want to map on to a genome (BAM, CRAM, BigWig, BigBed, indexed VCF, index GFF); JBrowse 2 generally does a pretty good job of guessing what sort of data file you want to add based on its extension.
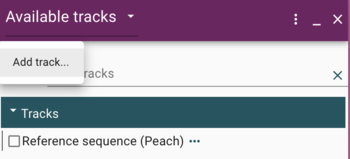
First, click one of the genome's "Open track selector" buttons; this will cause a new frame to open on the right side of the window, titled "Available tracks." Under that text is a "hamburger menu" icon (three horizontal lines). Click on that to get the "Add track" option.
We'll need the URLs for the data; in this case, we are using tabix-index GFF3. The URLs are:
Peach: https://s3.amazonaws.com/jbrowse.org/genomes/peach/Ppersica_298_v2.1.gene.sorted.gff3.gz https://s3.amazonaws.com/jbrowse.org/genomes/peach/Ppersica_298_v2.1.gene.sorted.gff3.gz.tbi
and
Grape: https://s3.amazonaws.com/jbrowse.org/genomes/grape/Vvinifera_457_v2.1.gene.sorted.gff.gz https://s3.amazonaws.com/jbrowse.org/genomes/grape/Vvinifera_457_v2.1.gene.sorted.gff.gz.tbi
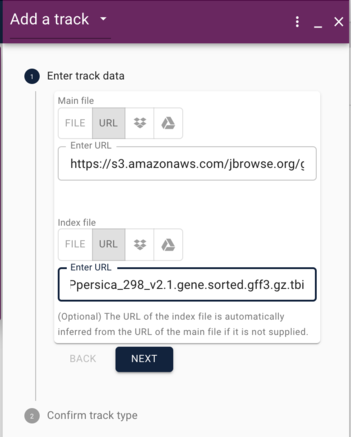
For which ever genome you are adding annotations, copy and paste the corresponding URL above into the "Main file" textfield, then copy and paste the tbi (index) file in the index file textfield. Then click the "Next" button.
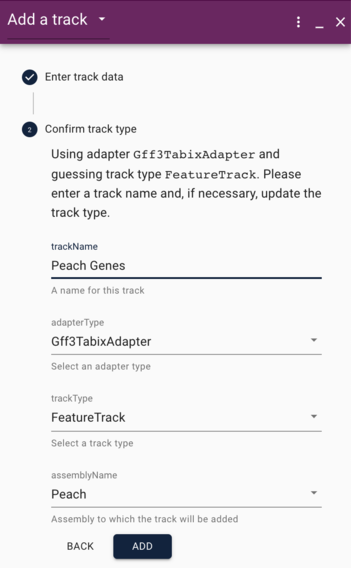
JBowse 2 will correctly guess that you are adding GFF3 data, so it will already have selected that option in the "Confirm track type" dialog, but one thing we will want to change here is the name of the track. JBrowse typically uses the name of the file to name the track, but file names are not always informative, so change the trackName entry to something useful like "Peach Genes" (unless of course, you're adding a Grape gene track). Also, double check that the "assemblyName" entry is what you expect. Now click the "Add" button, and repeat this whole procedure to add a gene track for the other species.
Depending on the zoom level of the synteny view, you will probably get a message about the gene tracks not getting loaded unless you zoom in or FORCE the loading of the tracks, which may make the application slow.
You can zoom in and out either using the magnifying glass icons in the upper right of each genome's frames or by clicking and dragging in the genome's "number line," or coordinate, region.
A note about preparing indexed GFF3 files
While we've provided prepared GFF3 files, if you wanted to do it yourself you would need GenomeTools (for sorting the GFF before indexing) and the bgzip and tabix packages that are part of Samtools. Sample commands are:
gt gff3 -sortlines -retainids -tidy yourfile.gff > yourfile.sorted.gff bgzip yourfile.sorted.gff tabix yourfile.sorted.gff.gz
Generally, you can navigate in the synteny view the way you would expect: by clicking and dragging anywhere in a genome's area other than the coordinate region (because clicking and dragging there will trigger the context menu that lets you zoom in). By default, this will cause only the genome that you're interacting with to move. This default can be changed by clicking on the "Toggle linked scrolls" icon in the upper left hand corner of the window (the oval with a line through it). Note that the other two icons next to the linked scroll icon don't actually do anything yet--we are planning implementation for those soon.
Getting data from other JBrowse instances
Getting a single NCList track
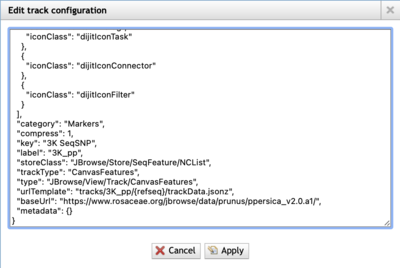
One under appreciated aspects of JBrowse is that it is quite open; if you can see a JBrowse page, you can pretty much always get at the underlying data. As an example of how this might work and be useful to you, we look at adding some SNP data for peach from the Genome Database for Rosaceae (GDR). The peach genome JBrowse that we want to look at is the one for the Prunus persica Genome v2.0.a1 assembly. This JBrowse 1 instance has several tracks, but we'll look at the 3K SeqSNP track. After opening that track, clicking on the down arrow in the label opens a menu and we want to look at "Edit config." This will open this dialog box, which you'll scroll until you find the urlTemplate entry:
The two pieces of useful information here are urlTemplate and baseUrl. We can combine those to make a full URL that we can use in our JBrowse Desktop application. In this case, just concatenating them will result in a URL that is very similar to the two for gene tracks that we used above:
https://www.rosaceae.org/jbrowse/data/prunus/ppersica_v2.0.a1/tracks/3K_pp/{refseq}/trackData.jsonz
Straight concatenation doesn't always work, but most of the time it does. If you are trying something like this, one thing you can do is test with a "real" chromosome name substituted in for the {refseq} part, like
https://www.rosaceae.org/jbrowse/data/prunus/ppersica_v2.0.a1/tracks/3K_pp/Pp04/trackData.jsonz
If clicking on that link gives you a 404, you did something wrong; if the browser asks to start a download, you did it right.
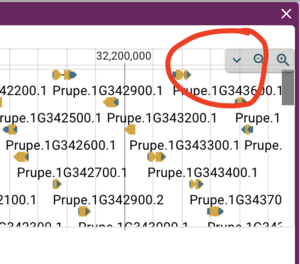
Now that we have an NCList url (the default track type for JBrowse 1--you can tell it's NCList from the file name, trackData.jsonz), we can do the same thing as before for adding the gene tracks. To make sure you have the correct "available tracks" window, click on the down arrow in the upper right hand corner of the peach genome frame, it looks like this:
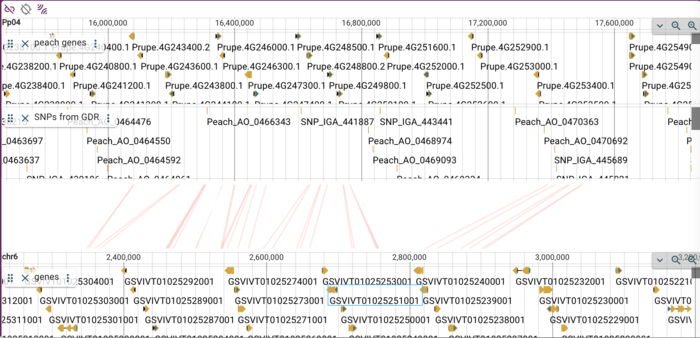
After opening the menu, select "Open track selector." and then proceed to add a new track just as before (click on the hamburger menu, then select "add new track" and go through the dialogs to add a new track using the first rosaceae URL above. Note that for NCList, there is no index file, so leave that field blank. Don't forget to change the track name to something useful! The result is a track that now has SNPs from GDR:
Possible point of failure: if you did everything right, you may still not have SNPs in your track. Check the track settings by clicking on the ... next to the track name and look at the URL for the NCListAdapter. Specifically, look for the curly braces { } in the url. If they were replaced with "% something something" they won't work, but putting the braces back will fix it.
Getting all of the tracks from a JBrowse 1 site
Add content here about getting peach jbrowse from https://www.rosaceae.org/jbrowse/data/prunus/ppersica_v2.0.a1/
Changing the way tracks look
JBrowse 2 gives users many ways to change the way tracks and the user interface look; here we'll look at a few examples.
Simple View Changes
Flipping the View
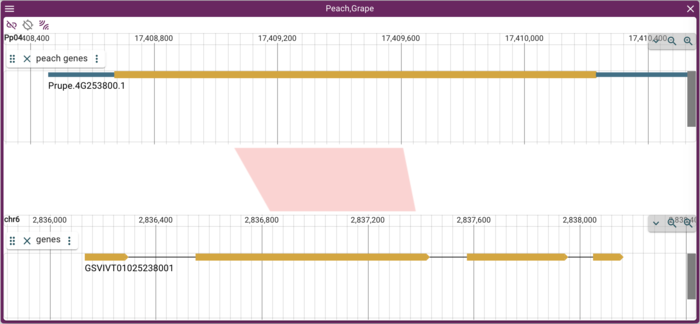
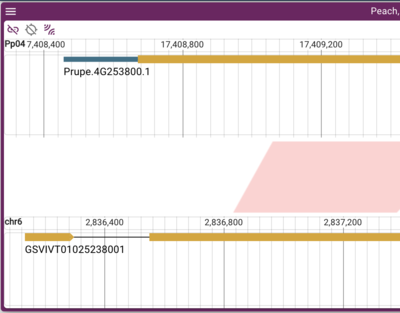
When doing work with synteny, it is frequently useful to be able to flip the direction that one of the genomes is displayed in, so that it can align with a syntenic region in the opposite strand of the compared genome. To see how this works, zoom into a single gene in one genome that has synteny in the other genome, and then zoom in to the related gene in the other genome (ie, so that there is only a single gene in each genome view). It will either look like this:
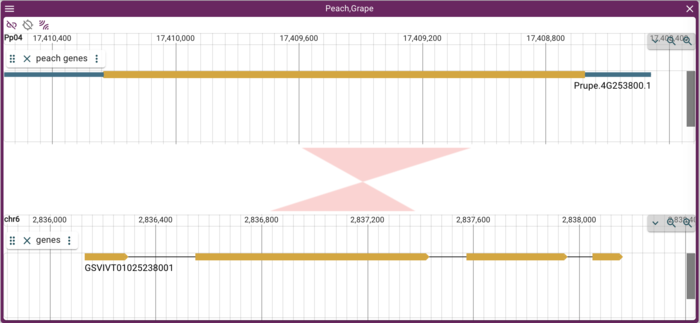
or like this:
You can switch between the "two triangles view" and the "trapezoid view" easily (terms that I literally just coined while writing this section). In the upper right corner of the synteny frame, there is a hamburger menu. When you click on that, you get options for the view 1 and view 2 menus (for each genome). Pick one and let the larger "per view" menu load. There are lots of options here, but the one we are interested in is "Horizontally flip." Selecting that will flip one genome and the shape of the synteny connector along with it.
Changing track label locations
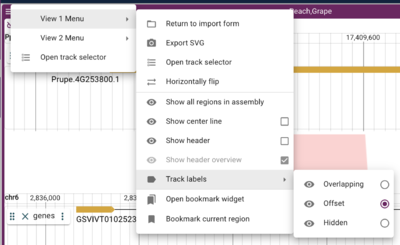
By default, the location of track labels in JBrowse is for them to overlap the contents of the track. This is because JBrowse views can get quite tall and this placement conserves height. The issue that users frequently have with this placement though is that it can obscure the features and labels that are placed under the translucent label, requiring them to pan to the left just to see the boundary of a feature or its label. JBrowse 2 gives two options for changing this default and both are accessed view the hamburger menu in the upper left corner of the synteny frame. Again, there are menus for each genome view; selecting one of those expands with multiple view options, one of which is "Track labels", which has three items as options:
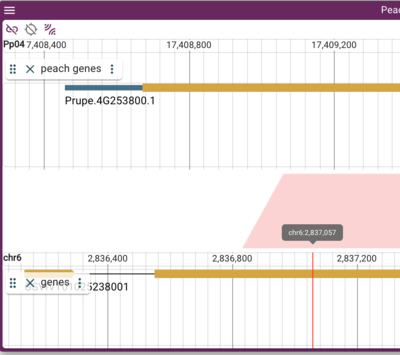
Selecting "Offset" puts the labels in their own vertical space making the display taller (much taller if you have multiple tracks open). Here is an example where the upper genome has offset labels and the lower genome has overlapping labels (note the obscured feature label):
and here is the same view with the track labels hidden. While hiding track labels may seem like an option you might not want, if you only have gene tracks in you synteny view, it would be "obvious" what the features are, so no labels would be needed.
Making an SVG of a genome
Several view types in JBrowse 2 support exporting of SVG images that are nice for using in publications. Unfortunately, at the moment, the synteny portion of the synteny view we've created does not support SVG output, but the linear genome view portions do support SVG output. To see an example of that, again open the hamburger menu in the upper left of the display and pick one of the two genome views. The second item in the view menu is "Export SVG". Selecting that will give you a dialog asking if you want to rasterize the canvas based tracks (which these are). I generally keep the default to rasterize, but you'll have to determine what is right for you given the intended purpose of the file. Here is an example SVG (not that it's real interesting):
Changing colors
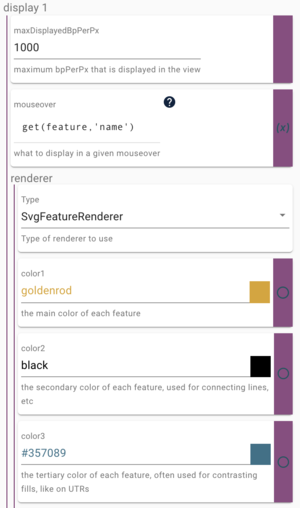
Finally in this section, we will change some aspects of how features are displayed in the track. You may have noticed that the default color for every feature is lovely goldenrod, a sort of dark yellow. It is NOT my favorite color. Fortunately, JBrowse makes it very easy for us to change the color of features. In this example, we change the color of the peach genes. There are really three colors we can change: the color of the CDS region (color1), the color of the intron connector (the thin black line, color2) and the color of the UTR region (color3). Since these are peach genes, we could try #ffe5b4 for the CDS color, which I would say is approximately a peach color. To edit the way the features in the track look, we need to have the "Available tracks" frame open. If it isn't already, in the upper right corner of the peach genome view, click on the "v" to open the context menu and select "Open track selector." Next to the peach genes track option, click on the "..." to open its context menu and select "Settings." There are quite a few options that can be adjusted in this control panel, but the one we are looking for is "color1" in the display1, renderer section:
Where it says "goldenrod" under color1, paste in "#ffe5b4" and the change will take effect immediately. While it was a cute idea to use a peach color for the CDS region, I think it is too light, so lets pick another color. The color box next to the color name (which right now is a peach color) is actually a button to bring up a color picker. Pick a color you like, and again, when you pick a color, the change happens immediately. You can do the same for the color2 and color3.
Using JavaScript/JEXL to code changes
This is a slightly more advanced topic. In addition to changing track settings by changing the names of colors, we can also change aspects of the how the track looks and behaves by adding snippets of JavaScript or a JavaScript-related language called JEXL referred to as callbacks. We'll look at two examples here to give a flavor of the sorts of things you can do.
Changing the color of a glyph according to strand
Since we were looking at glyph color in the previous section, we'll stay there and make the glyphs color change according to the strand that the gene is on. First note that on the right hand side of most of the fields in the track settings dialog is a circle in the purple box. When you mouse over that circle, the mouse hover text says "Convert to callback." First check that circle for the "color1" field. When you do that, you may notice that the gene glyphs in the track turned black--that's because JBrowse expects there to be a snippet of code there, and there isn't, so you get the "I don't know what to do" color, which is black.
Next we'll add the code here to the color1 field:
get(feature, 'strand')>0?'blue':'red'
What it is doing is quite simple: it says get the feature's strand and if it's positive, make the feature blue, otherwise make it red. Once again, you should see an immediate change in the way the track looks:
Changing the hover text
Another example that is pretty easy is modifying the text that appears in the mouse over hover box. The default in JBrowse is generally the name of the feature, and it is already a callback (i.e., we don't need to check the circle to make it one). That callback looks like this:
get(feature,'name')
Now we want to look for something useful to add. Frequently in GFF files, there is extra information in the ninth column that users might want to see. The the peach and grape GFF files, there wasn't much extra information, but we'll make do. The peach gene annotations have an attribute called "longest." For a given gene, the transcript that is the longest has this attribute set to 1, the rest are zero. What we will do is add "longest transcript" to the mouse over when that's true (and of course, when there is only one transcript for a gene, what will happen to the mouse over?). To do this, we can modify the original callback to look like this:
get(feature,'name')+(get(feature,'longest')>0?' longest transcript':' ')
This is very similar to the strand callback: it's saying get the feature name and concatenate it (with the "+" operator) with different text depending on the result of the question of whether the "longest" value is greater than zero.
Using JBrowse server
The server version of JBrowse 2 is set up in a way that is very similar to this. To install, the prerequisites are quite simple:
- a web server like Apache or Nginx
- NodeJS version 10 or better
That's really it for the server. Other things the would likely help include GenomeTools for sorting GFF, and Samtools for working with BAM and CRAM files, which also provides tabix which is used for indexing GFF and VCF files.
JBrowse admin
JBrowse 2 also has an admin server that has a user interface for editing the configuration that works very much like the JBrowse desktop version we've been using in this tutorial. For more information on it, see the JBrowse website https://jbrowse.org/jb2/docs/quickstart_gui/.