NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
GMOD Online Training 2014//SOBA Tutorial
Contents
About SOBA
SOBA is a command line tool and web application for analyzing GFF3 annotations. GFF3 is a standard file format for genomic annotation data. SOBA gathers statistics from GFF3 files and renders them as tables and graphs.
The web version of SOBA will produce the following:
- Summary statistics of feature types and attributes used
- Histograms of feature lengths
- Graphs of Sequence Ontology terms used
- Histograms of intron density
- Suggestions to improve SO compliance for invalid terms
In addition, the command line tool (SOBAcl) flexibly produces a much wider variety of tables, figures and graphs based on the data in a GFF3 file, as well as the ability to produce complex and extensible custom reports via a robust template system.
SOBA is a tool for those dealing with genomic sequence annotation who want to view genome-wide summaries of their annotation files. For example: SOBA would be a useful tool at an annotation jamboree for a newly sequenced organism and when preparing the resulting genome paper; SOBA would help those developing annotation tools to quickly evaluate updates to their tool; SOBA assists comparative genomics analyses by providing a high-level overview of the genome of multiple organisms. SOBA complements genome browsers by providing a summary of all the features annotated in the genome.
SOBA is built with Perl (and JavaScript for the web interface). The web interface uses CGI::Application as a Perl webapp framework and the JQuery JavaScript library for Web 2.0 effects and AJAX. Both versions of SOBA use the Template Tooklit (TT) to generate html/txt reports, graphviz for the ontology graphs, and GD for charts. Template Toolkit makes extensibility very easy, at least for someone who's willing to learn the fairly simple template language of TT as you don't need to know Perl or any other programming to use TT.
SOBA Web Application
Documentation for the web interface to SOBA is available on the Sequence Ontology Wiki as well as via tool-tips on the site itself.
Navigate to the SOBA Web Application with any modern browser that has JavaScript enabled.
SOBAcl
Information and downloads for SOBAcl are available from the Sequence Ontology website.
Tutorial AMI
- NAME: GMOD Summer 2014 - SOBA/GAL
- ID: ami-58846d30
Getting Started
Let's start by grabbing some data to play with.
cd /home/ubuntu/ # wget topaz.genetics.utah.edu/SOBA_GMOD_demo.tar.gz # tar -xzvf SOBA_GMOD_demo.tar.gz # cd soba/ # export PATH=$PATH:/usr/local/SOBA/bin #The above steps are already set up on the AMI cd soba_examples # The GO OBO Parser is missing on this image - but we don't need it today. # To install it for you own use later do: # sudo cpan GO::Parser
Help
Documentation for the command line version, SOBAcl, is available as a usage statement with the script itself:
SOBAcl --help
Synopsis:
SOBAcl [options] db_name
SOBAcl [options] file.gff3
SOBAcl file.gff3 2> soba_count.txt
Description:
The Sequence Ontology Bioinformatics Analysis command line tool
(SOBAcl) will generate a variety of reports from the data in GFF3
files. The data can optionally first be loaded into a database.
Options:
--help Print this help page and exit.
--title A title for the table, graph, or report.
--columns [--series]
The columns parameter indicates how data will be
grouped into columns (or series for axis graphs and
seperate graphs for pie graphs). For example, using
seqid as the value would create one column of data for
each seqid (chromosome). For bar and line graphs there
would be a data series for each value. For pie graphs,
a separate graph would be generated for each
value. (seqid, source, type, strand, phase, [file],
stats).
--rows [--x_data]
The rows parameter indicates data will be grouped into
rows (x-axis labels for graphs with axes; wedges for
pie graphs). For example, using type as the value
would create one row of data for each unique feature
type in the GFF3 file. For bar and line graphs there
would be one x-axis label for each type and for pie
graphs there would be one pie wedge for each
type. (seqid, source, [type], strand, phase, file)
--data [--y_data]
The data parameter indicates which data fields (columns
or attributes in the GFF3 file) will be reported. For
example using score for the data parameter will create
a report on the scores of features. (seqid, source,
type, start, end, [length], score, strand, phase)
--data_type The data_type parameter indicates how the data described
in the data parameter should be summarized. For
example if data is length and data_type is 'mean', then
the mean length would be reported for each grouping of
the data. (count, [mean], harmonic_mean,
geometric_mead, median, mode, sum, min, max, variance,
standard deviation, range, footprint)
--layout The layout parameter describes how the data will be
presented. ([table], lines, bars, hbars, points,
linespoints, area, pie)
--format The format parameter determines how the output will be
formatted. For all text based outputs the options are
(text, [tab], html, html_page, pdf). For all graphical
layouts the options are (jpeg, [png], gif).
--collapse Collapse series data onto one graph.
Tables
First let's just run SOBAcl with all default parameters and see what we get...
SOBAcl hsap_hg18_demo.gff3
By default SOBA prints count of each feature type.
type type (count)
=====================================
| |hsap_hg18_demo.gff3|
=====================================
|CDS |11944 |
+---------------+-------------------+
|contig | 3 |
+---------------+-------------------+
|exon |12918 |
+---------------+-------------------+
|five_prime_UTR | 1381 |
+---------------+-------------------+
|gene | 1157 |
+---------------+-------------------+
|mRNA | 1085 |
+---------------+-------------------+
|ncRNA | 72 |
+---------------+-------------------+
|three_prime_UTR| 1385 |
-------------------------------------
Next let's customize the report by specifying the data that we want to summarize in the table rows.
SOBAcl --rows seqid hsap_hg18_demo.gff3
seqid seqid (count)
===========================
|seqid|hsap_hg18_demo.gff3|
===========================
|chr1 |9986 |
+-----+-------------------+
|chr2 |9977 |
+-----+-------------------+
|chr3 |9982 |
---------------------------
Now we will further customize the table by indicating what data we want to represent in our rows and columns.
SOBAcl --rows type --columns seqid hsap_hg18_demo.gff3
type type (count)
================================
|type |chr1|chr2|chr3|
================================
|CDS |3815|4144|3985|
+---------------+----+----+----+
|contig | 1| 1| 1|
+---------------+----+----+----+
|exon |4182|4416|4320|
+---------------+----+----+----+
|five_prime_UTR | 540| 387| 454|
+---------------+----+----+----+
|gene | 463| 318| 376|
+---------------+----+----+----+
|mRNA | 430| 301| 354|
+---------------+----+----+----+
|ncRNA | 33| 17| 22|
+---------------+----+----+----+
|three_prime_UTR| 522| 393| 470|
--------------------------------
By default we've just been getting the counts of the different feature types. Let's look at their lengths by specifying length to the --data parameter and mean to the --data_type parameter.
SOBAcl --rows type --columns seqid --data length --data_type mean hsap_hg18_demo.gff3
type length (mean)
========================================================
|type |chr1 |chr2 |chr3 |
========================================================
|CDS | 168.67| 169.28| 159.59|
+---------------+------------+------------+------------+
|contig |247249718 |242951148 |199501826 |
+---------------+------------+------------+------------+
|exon | 308.16| 270.85| 287.11|
+---------------+------------+------------+------------+
|five_prime_UTR | 554.27| 618.66| 629.51|
+---------------+------------+------------+------------+
|gene | 48070.89| 88243.71| 73324.19|
+---------------+------------+------------+------------+
|mRNA | 50172.13| 90551.96| 77185.4 |
+---------------+------------+------------+------------+
|ncRNA | 20691.15| 47374.06| 11193.95|
+---------------+------------+------------+------------+
|three_prime_UTR| 539.2 | 579.47| 596.2 |
--------------------------------------------------------
Let's look at the minimum feature length for the entire file.
SOBAcl --rows type --data length --data_type min hsap_hg18_demo.gff3
type length (min)
=====================================
|type |hsap_hg18_demo.gff3|
=====================================
|CDS | 2 |
+---------------+-------------------+
|contig |199501826 |
+---------------+-------------------+
|exon | 6 |
+---------------+-------------------+
|five_prime_UTR | 0 |
+---------------+-------------------+
|gene | 74 |
+---------------+-------------------+
|mRNA | 287 |
+---------------+-------------------+
|ncRNA | 74 |
+---------------+-------------------+
|three_prime_UTR| 0 |
-------------------------------------
Specifying --data_type footprint will give us a look at the "tiled" length of the features on the chromosome.
SOBAcl --rows type --data length --data_type footprint hsap_hg18_demo.gff3
type length (footprint)
=====================================
|type |hsap_hg18_demo.gff3|
=====================================
|CDS | 1829153 |
+---------------+-------------------+
|contig |247249719 |
+---------------+-------------------+
|exon | 3423479 |
+---------------+-------------------+
|five_prime_UTR | 785362 |
+---------------+-------------------+
|gene | 61344223 |
+---------------+-------------------+
|mRNA | 60303556 |
+---------------+-------------------+
|ncRNA | 1627355 |
+---------------+-------------------+
|three_prime_UTR| 717369 |
-------------------------------------
Perhaps we want to load that data into Excel so the we can prepare a table or figure for our paper.
SOBAcl --rows type --data length --data_type footprint --format tab hsap_hg18_demo.gff3
hsap_hg18_demo.gff3 CDS 1829153 contig 247249719 exon 3423479 five_prime_UTR 785362 gene 61344223 mRNA 60303556 ncRNA 1627355 three_prime_UTR 717369
Charts
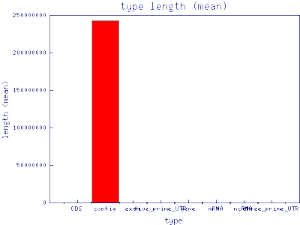
Let's start by looking at the mean length of the features.
SOBAcl --x_data type --series seqid --data length --data_type mean \ --layout bars --format png hsap_hg18_demo.gff3 firefox SOBAcl_graphic_chr*.png
That worked, but it doesn't look quite like what we had in mind. First of all, the X-axis labels overrun each other, the contig feature is so long that none of the other feature lengths even show up on the chart, and we've got three charts - one for each chromosome (seqid). Let's fix all of those problems using the --gd, --collapse and --select parameters.
SOBAcl --x_data type --series seqid --data length --data_type mean \ --layout bars --format png --gd x_labels_vertical=1 --select 'type => "exon"' \ --collapse hsap_hg18_demo.gff3 firefox SOBAcl_graphic.png
If we just wanted to exclude the contigs, our --select parameter can do that as well.
SOBAcl --x_data type --series seqid --data length --data_type mean \ --layout bars --format png --gd x_labels_vertical=1 --select \ 'type => ["ne", "contig"]' --collapse hsap_hg18_demo.gff3
We can constrain the features reported in other ways as well.
SOBAcl --columns seqid --rows type --data length --data_type mean \ --layout table --format text --select 'start => [">=", "1000"], \ end => ["<=", "1000000"]' hsap_hg18_demo.gff3
Reports
SOBAcl has support for more complex reports.
SOBAcl --report attributes --format html_page hsap_hg18_demo.gff3
These reports can be generated by custom templates.
SOBAcl --columns file --rows type --data length --data_type mean \ --layout table --format tab --template soba_custom_template_text.tt \ hsap_hg18_demo.gff3
count length (mean) CDS 11944 165.853064300067 contig 3 229900897.333333 exon 12918 288.366697631212 five_prime_UTR 1381 597.052136133237 gene 1157 67319.117545376 mRNA 1085 70187.8202764977 ncRNA 72 24089.3611111111 three_prime_UTR 1385 569.969675090253
---
Genome Annotation Library
The Genome Annotation Library (GAL) is a library of Perl modules that makes working with sequence features in GFF3 format straight forward and intuitive. It aims, 'to make easy things easy and hard things possible', when working with GFF3 based sequence feature data. GAL also provides numerous parser classes for converting other formats to GFF3 and makes writing new parsers easy. The GAL API takes advantage of the powerful ontology based relationships copatured by GFF3-based sequence feature data to allow users to traverse the relationships between features in a GFF3 file. GAL loads features into a light-weight relational database and thus selecting sets of features based on complex data queries (get all genes on the minus strand of chromosome 2 who have a Dbxref attribute set) is straight-forward, and iterator classes make working with feature sets convenient.
In addition to the GAL programming library many scripts utilizing the library are provided as well as several stand-alone scripts for working with GFF/GVF, fasta, fastq, and SAM/BAM files are included (see below).
GAL is written in Perl and uses DBIx::Class to interface with an SQLite representation of the sequence features. See the Documentation link below to find out more.
Stand-alone Scripts
Several popular stand-alone scripts (they don't use the underlying GAL library) are included with GAL and can be used without installing the GAL libary.
- fasta_tool
- A script providing various utility functions to manipulate fasta files.
- gff_tool
- A script providing various utility functions to manipulate GFF files.
- gtf2gff3
- A script to convert GTF to GFF3 files.
- gff3_validator
- A syntactic validator for GFF3 files (Does not validate relationships).
- gvf_validator
- A syntactic validator for GVF files.
- ucsc2gff
- A script to convert USCS gene tables to GFF3 output. Works with the build_genes script below.
- build_genes
- A script to add genes to UCSC table based GFF3 files. Works with ucsc2gff above.
If you are only intersted in using these scripts they can be downloaded as GAL_#.#.#_stand_alone_scripts.tar.gz from the same location described in the Download section.
Download
GAL releases can be downloaded from:
http://www.sequenceontology.org/software/GAL_Code/
The bleeding-edge code for GAL is also available (via Subversion) from:
svn co svn://topaz.genetics.utah.edu/GAL/trunk GAL
Installation Installation is automated by the included installation script and an INSTALL document describes all the details.
Documentation
Full API documentation can be found with the distribution in the [GAL/docs/html/GAL.html GAL/docs/html/GAL.html] document or online at:
GAL Manual
In addtion, each script included with GAL comes with a detailed useage statement - just run the script with the --help flag.
GAL Working Example
A very simple working GAL script below will iterate over mRNA features, retrieve exons, infer and retrieve introns and call methods on each resulting object.
use GAL::Annotation;
my $annotation = GAL::Annotation->new( $gff3_file, $fasta_file );
my ( $gff3, $fasta_file ) = @ARGV;
my $features = $annotation->features;
# Do the search and get an interator for all matching features.
# Searches can be as complex as desired.
my $mrnas = $features->search( { type => 'mRNA' } );
# Iterate over the features
while ( my $mrna = $mrnas->next ) {
# Get the feature ID
my $mrna_id = $mrna->feature_id;
# Get and iterator for all the exons for this mRNA
my $exons = $mrna->exons;
# The iterators are smart
my $e_count = $exons->count;
my ( $e_length, $e_gc );
# Iterate over each exon
while ( my $exon = $exons->next ) {
$e_length += $exon->length;
$e_gc += $exon->gc_content;
}
print join "\t", int( $e_length / $e_count ), $e_gc / $e_count;
print "\n";
# Introns don't exist in the dataset, so GAL
# will infer them on the fly.
my $introns = $mrna->introns;
my $i_count = $introns->count;
my ( $i_length, $i_gc );
while ( my $intron = $introns->next ) {
$i_length += $intron->length;
$i_gc += $intron->gc_content;
}
print $i_length
? join( "\t", int( $i_length / $i_count ), '', $i_gc / $i_count )
: "\t\t";
print "\n";
}
Mailing List GAL is supported by the Sequence Ontology Developers Mailing list at: